Monte-Carlo sampling is used toĬalculate p-values for all edges in the tree. Other.Afterwards, a minimum spanning tree is built from rescaled node coordinates. These nodes are further distorted in 2D space based on their density of similarity to each That dimensionally reduced spaced is compresed into a 2D representation of similarities between neighboring nodesĪcross the SOM network. Unsupervised training of the SOM produces a low-dimensional reprentation of input space. The AutoSOME algorithm revolves around the use of a Self-Organizing The normal visualization is to create a new networkĪutoSOME clustering is the one cluster algorithm that functions both as an attribute cluster algorithmĪs well as a network cluster algorithm. network clustering: an edge attribute is chosen to partition the network.Selection of cell in the heat map selects an edge in the network. If an edge attribute is chosen, the columns also correspond to the nodes in the network and The normal visualization is some kind of heat map where the rows correspond to the nodes in the network. attribute clustering: a list of node attributes or a single edge attribute is chosen to perform the clustering.There are two different types of clustering algorithms supported by clusterMaker: Will be available in a session that was saved after clustering. Hierarchical or k-Means clusters if either of those methods had beenīecause information about clusters is saved inĬytoscape attributes, the Eisen TreeView and Eisen KnnView options (without clustering), and options appropriate for displaying This will bring up the settings dialog for the selected algorithmĪ subset of the visualization options, including showing a heat map of the data Where algorithm is the clustering algorithm you wish to use (see Figure 2).
New Cluster menu hierarchy under the Plugins main menu.Įach of the supported clustering algorithms appears as a separate
#Clustering app cytoscape install
Once clusterMaker is installed, it will install a Select clusterMaker and click the Install button.įigure 2. Select Analysis under Available for Install. Is available in the Analysis group of plugins. You must be running Cytoscape 2.8.2 or newer.
#Clustering app cytoscape download
To download clusterMaker using the plugin manager,
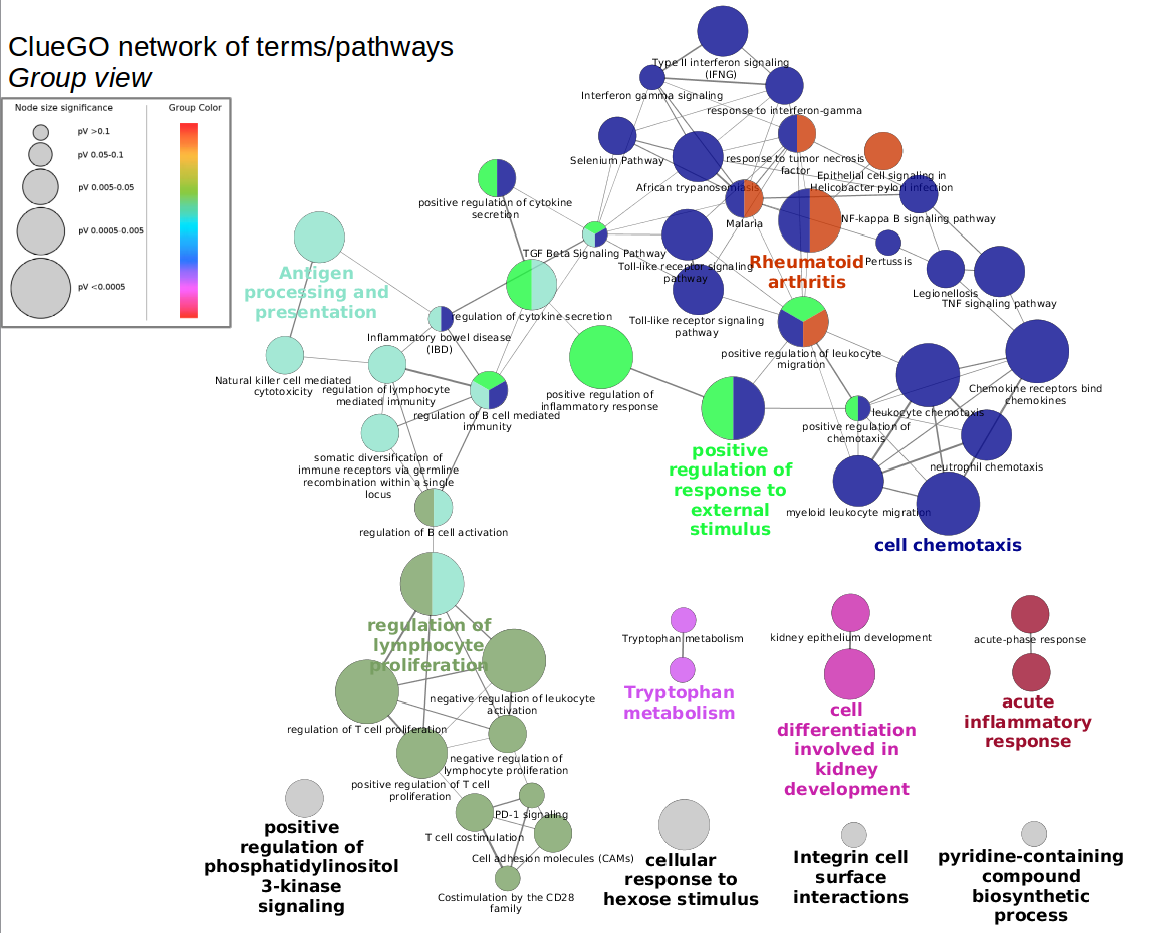
In this screenshot, theĮxpression data in the sampleData file galFiltered.cys has been clustered using the hierarchical methodĪnd displayed as a heatmap with associated dendrogram.

